RNA Frabase 2.0
The search engine for RNA fragments and structures. Find your favorite particle today!
Technologies: #Linux #PostgreSQL #AWK #Apache #PHP #HTML #CSS #JavaScript #LAMP
About
How to transform scientific theory into a website? With some serious problem solving, obviously. I was responsible for implementation, cloud deployment and optimization of the search engine. The website was a success and is still widely used in the research community (2,8 mln queries). You check it out here: http://rnafrabase.cs.put.poznan.pl .
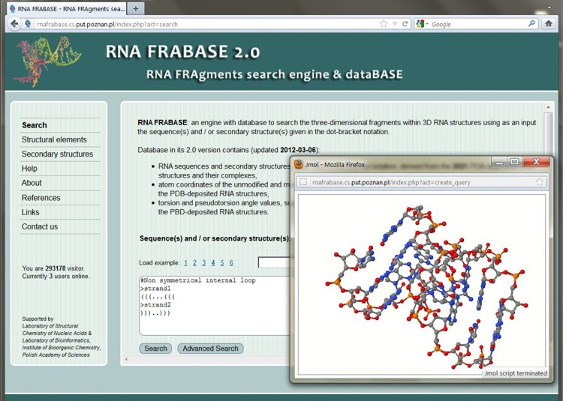
Knowledge of the three dimensional RNA structure is crucial for all fields of biomolecular research. In contrast to the protein field, only about 1.300 experimentally derived structures of RNAs are deposited in the Protein Data Bank (PDB). To complement the results of experimental studies, new approaches based on bioinformatics and calculation are pursued in several laboratories to make tertiary RNA structure prediction possible. Here, we present the RNA FRABASE version 2.0, which should greatly facilitate various RNA structure modelling approaches, RNA structure analysis and motif searching.
If one compares the three dimensional RNA structure to a spatial puzzle, the RNA FRABASE allows to pull out a defined piece of this puzzle - the 3D RNA fragment. The architecture of the web-accessible RNA FRABASE engine and database is based on the following information path: PDB-deposited RNA structures » RNA sequences and secondary structures described in the dot-bracket notation » secondary structures of RNA fragments » 3D RNA fragments.
In the 2.0 ver. database contains RNA sequences and secondary structures, described in the dot-bracket notation, derived from 2753 PDB-deposited RNA structures and their complexes. It also contains atom coordinates of the unmodified and modified nucleotide and nucleoside residues (1921887 cases) extracted from the PDB-deposited RNA structures, as well as torsion and pseudotorsion angle values, sugar pucker parameters and classification of base pair types. Also new functionalities “Structural elements” and “Secondary structures” were added.
Find your favorite RNA particle today!